Product Specifications
| Yield | Minimum of 200 ng of amplified dsDNA |
| Turnaround time | 8-12 business days |
| Delivery Format | Dried-down, dsDNA pooled in a 2 mL tube |
| Length | 301 - 500 bp |
| Pool size | No minimum, no maximum |
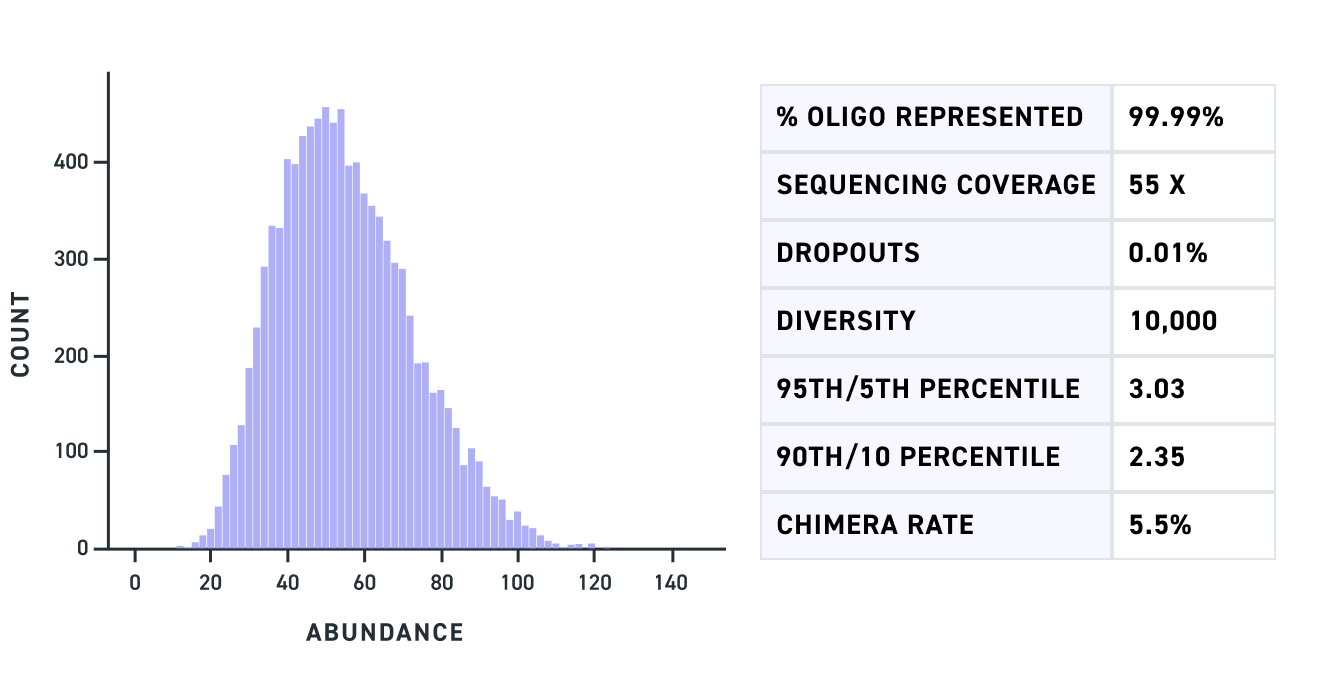
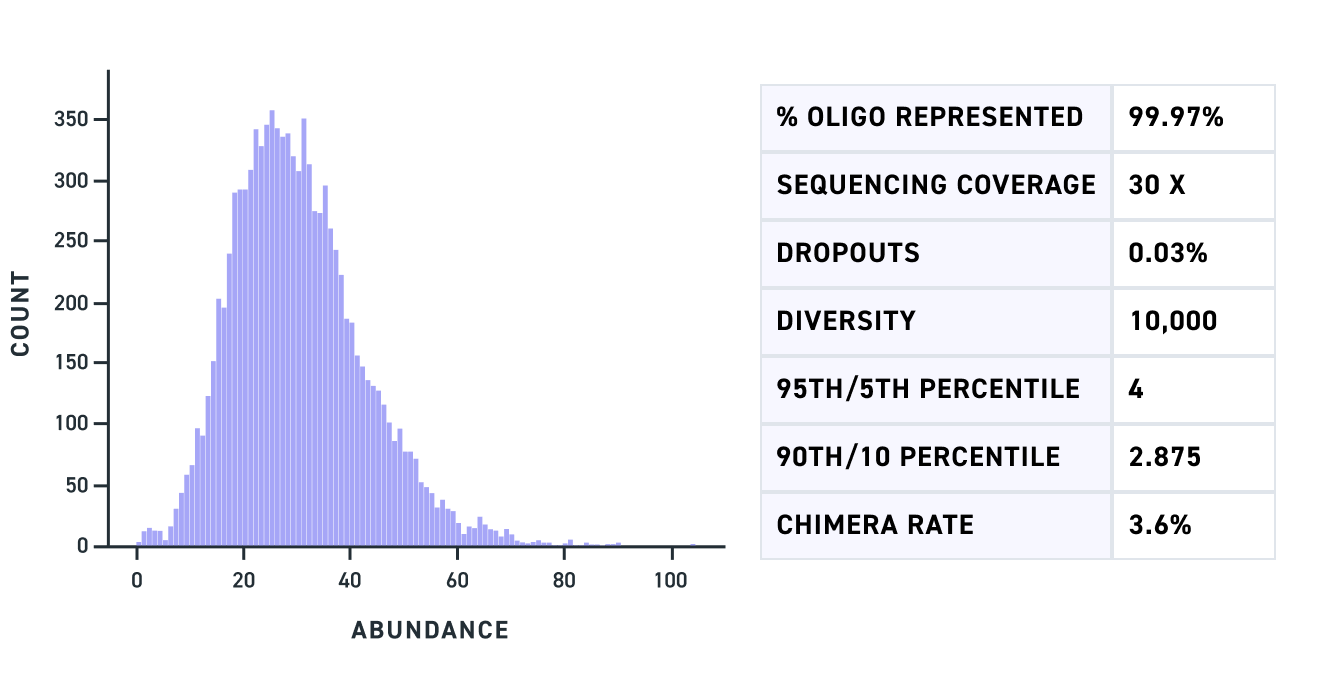
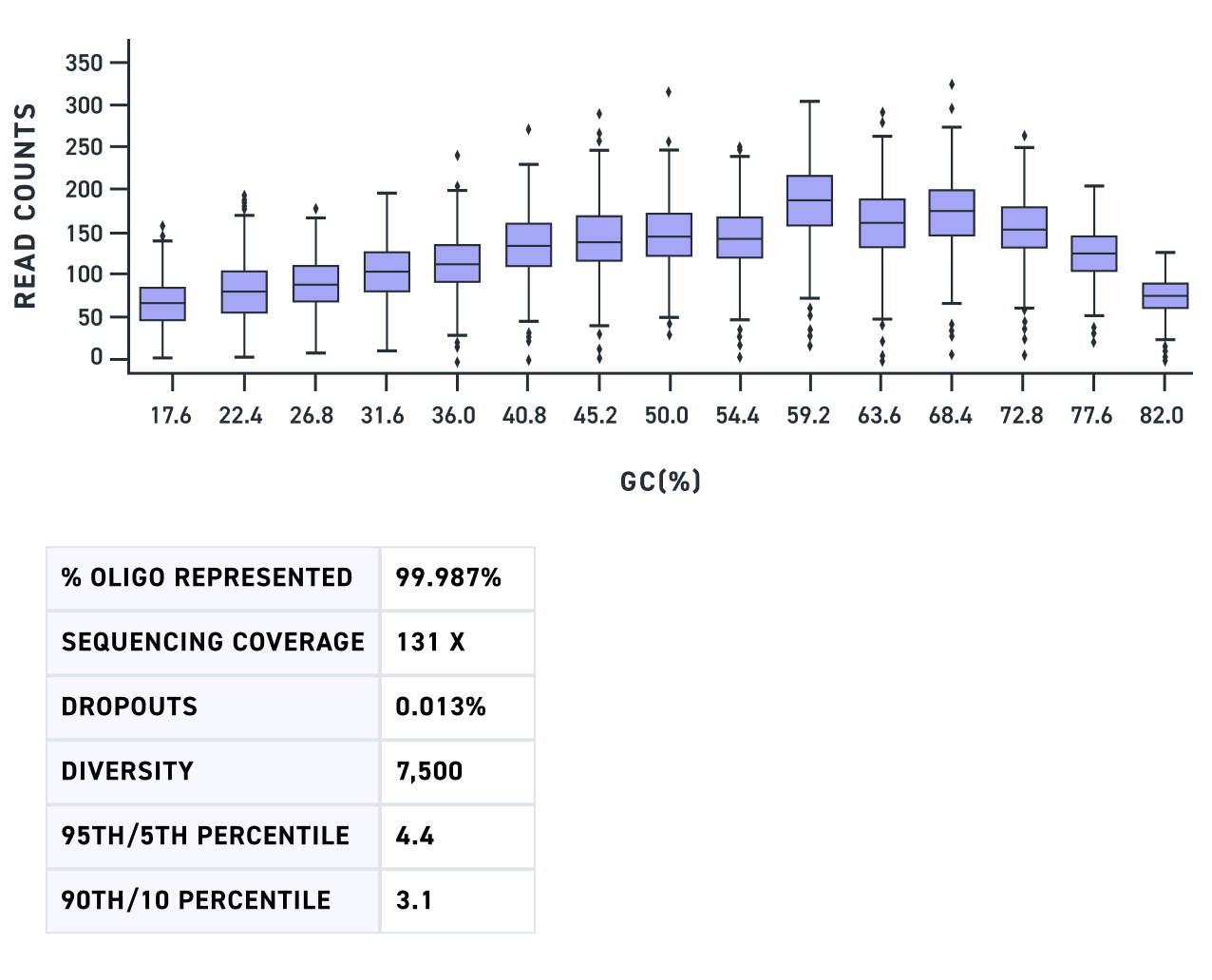
| Uniformity | 90% of sequences are within ~3x of the mean* |
| Quality Control (QC) | Fragment analysis to ensure >90% of genes in a pool are the correct length* |
| Error Rate | 1:2000 nt (from the original pool synthesis, before amplification) |
*Terms and Conditions: Standard turnaround time for Multiplexed Gene Fragments is 8-12 business days. This will vary based on sequence length and complexity. Multiplexed Gene Fragment length is up to 500 base pairs, with up to 460 bp allotted for the variable region with a minimum of 40 bp conserved for the primer region. Quality control tests ensure >90% of gene sequences within a pool will be user defined length.
For research use only. Not for Diagnostic Procedures.









.png)